Esha Manchanda

Hi! I am Esha, a Masters in Biomedical Data Science student at Nanyang Technological University, Singapore. I graduated from Ashoka University with a Bachelors in Computer Science. I am passionate about improving human health through the confluence of technology and personalised medicine. My research interests lie at the intersection of neuroscience, immunology, artificial intelligence and epigenetics. I am especially interested in reversing neurodegeneration and remyelination in multiple sclerosis.
I have leveraged computational modeling, data science and machine learning across various healthcare projects. At present, I am doing research at the Lo Lab at LKC Medicine.
Outside of academia I love playing badminton, running, solo travelling and learning korean!
Ongoing Research
-
Dysregulation of the Metabolic-Inflammatory Axis in Progressive Multiple Sclerosis
Esha Manchanda, Eka Saipuljumri, Jialiu Zeng, Chih Hung Lo
In Preparation
Presented as a talk and poster at SSBBI & Neuroscience Singapore 2024
SlidesMetabolic impairments are shown to be implicated in the pathogenesis of neuroinflammatory diseases such as MS, and therapies aimed at enhancing metabolic functions could potentially attenuate neuroinflammation and open new avenues for treatment of progressive MS. In this study, we are investigating the role of mitochondrial, autophagic, and lysosomal dysfunction in progressive MS, exploring their interconnected roles in driving neuroinflammation and neurodegeneration.
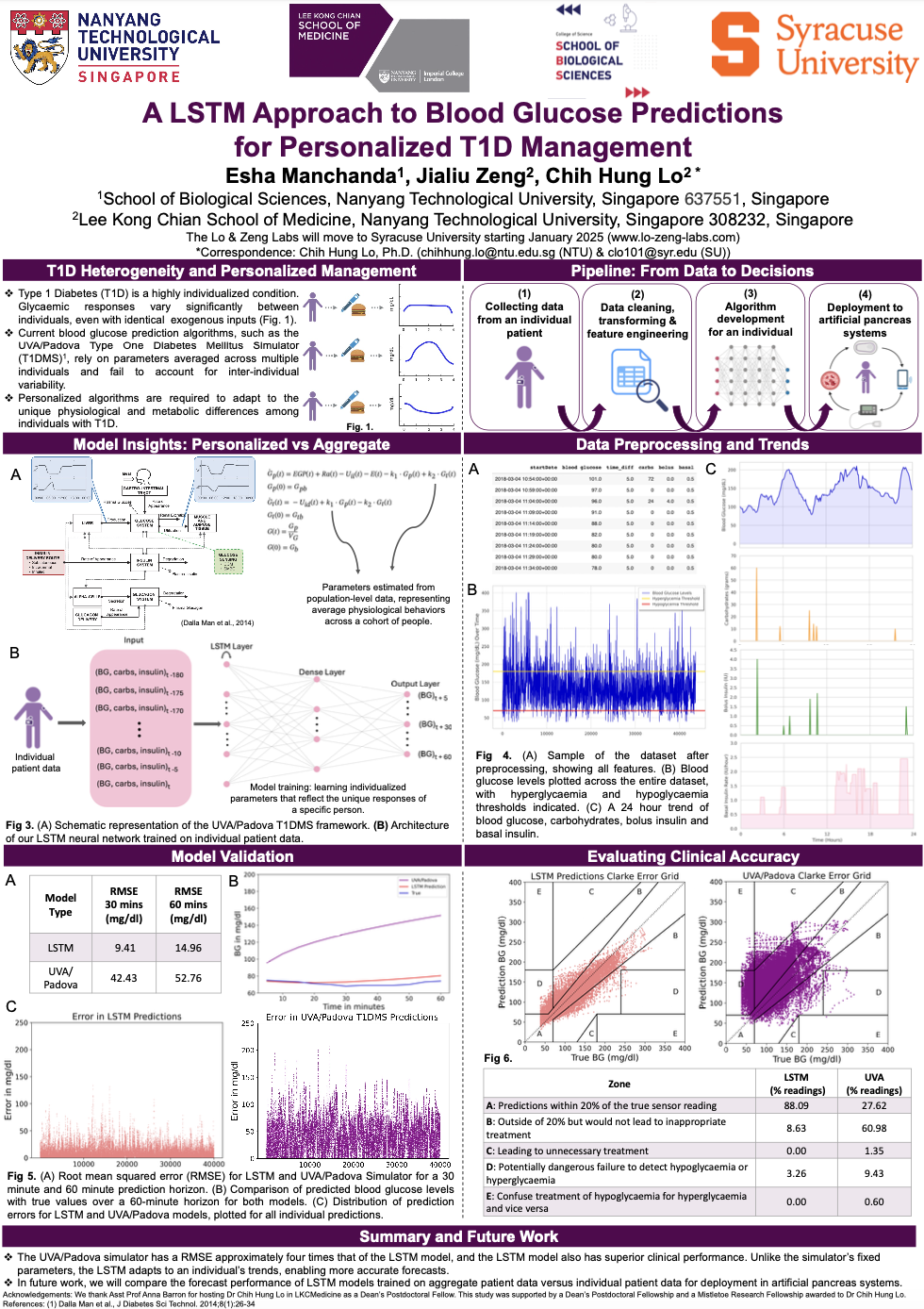
A LSTM Approach to Blood Glucose Predictions for Personalized T1D ManagementEsha Manchanda, Jialiu Zeng, Chih Hung Lo
In Preparation
Presented as a poster at FEBS-IUBMB-ENABLE 2024Type one diabetics need to be their own pancreas – it’s almost as though pumping blood to your body required having to know when it’s time to pump blood, rather than having your heart do it automatically. To ease the burden on T1Ds and improve blood glucose regulation, automated insulin delivery systems are being actively developed. Managing T1D, however, is significantly individualized— two people can have completely different blood glucose dynamics under identical exogenous conditions. Therefore, current systems are not well-suited to capture the unique disease patterns of a single individual. In this work, we have built a personalised blood glucose prediction model to address this inherent patient-patient variability.
Multiplexed Imaging as an Innovative Approach for Single-Cell Spatial Profiling of Brain PathologyEka Saipuljumri, Esha Manchanda, Jialiu Zeng, Chih Hung Lo
In Preparation
An important aspect of understanding neuroinflammatory and neurodegenerative pathologies lies in defining cell populations and their spatial interactions, as well as the environmental cues that drive their specific expression profiles in specific regions of the brain. While single-cell RNA-seq is superior in isolated cells or nuclei compared to tissue with regard to gene coverage and sequencing depth, highly multiplexed protein imaging has a parametric depth of only tens to hundreds of markers but allows for precise outlining of cells with complex and irregular morphologies, which will improve the spatial analysis of individual cells. In this study, we present an innovative approach through serial immunofluorescence staining of tissue sections with repetitive cycles of immunolabeling, scanning, and antibody removal, termed iterative indirect immunofluorescent imaging (4i) for single-cell spatial profiling of brain pathologies.
Past Work
-
Undergraduate Thesis Candidate, Ashoka UniversityAugust 2023 - May 2024
In my thesis, I explore the relationship between the Epstein-Barr Virus and Multiple Sclerosis. Through a mathematical modeling approach, my work studies the potential link between EBV and MS, closely examining the immune system's response to EBV infection. It investigates the role of T-cell exhaustion in uncontrolled EBV lytic activity in B cells, leading to chronic inflammation and potentially the development of MS.
Research Intern, Max Hospitals & Ashoka UniversityJanuary 2023 - June 2023Over the past year, I have developed a growing interest in the gut microbiome and its impact on health. I worked on a project employing model-agnostic meta-learning techniques aimed at predicting diseases through the targeted analysis of fecal gut microbiome data. Beyond this, my interest extends to exploring the intricate connections between the gut microbiome and various health dimensions, including its role in both wellness and disease pathology.
Computational Genomics Research Intern, Ashoka UniversitySeptember 2022 - December 2022Traditional bulk methods (techniques used to analyze biological samples that contain a large population of cells) analyze an average over many cells, which can mask variations between individual cells. Single cell analysis helps investigate the heterogeneity between cells. Deciphering cell-cell heterogeneity at the chromatin level is very important while developing accurate disease models and determining patient responses to specific drugs. I conducted a comprehensive review of algorithms for single-cell analysis of accessible chromatin (scATAC-seq) to assess the computational challenges associated with scATAC-seq analysis including inherently sparse data, determination of features, peak calling, the impact of sequencing coverage and noise, and clustering performance.
Teaching Experience
I really enjoy teaching, and inspired by my many great teachers, I adopt a constructivist teaching style—where I focus on guiding students to construct knowledge rather than passively absorb information.
Teaching Assistantships
-
Computational/Mathematical Biology (Spring 2024), Ashoka University
Taught by Professor Sudipta Tung
-
Introduction to Computer Programming (Spring 2022), Ashoka University
Taught by Professor Subhashis Banerjee | Course Content -
Computational/Mathematical Biology (Monsoon 2021), Ashoka University
Taught by Professor Sudipta Tung | Course Website
Workshops Conducted
-
IEEE Ashoka PySci Computational Biology Workshop (November 2023)
[Hands-on exercise] -
Computer Science Society Tools for Web Scraping (October 2021)
[Code]